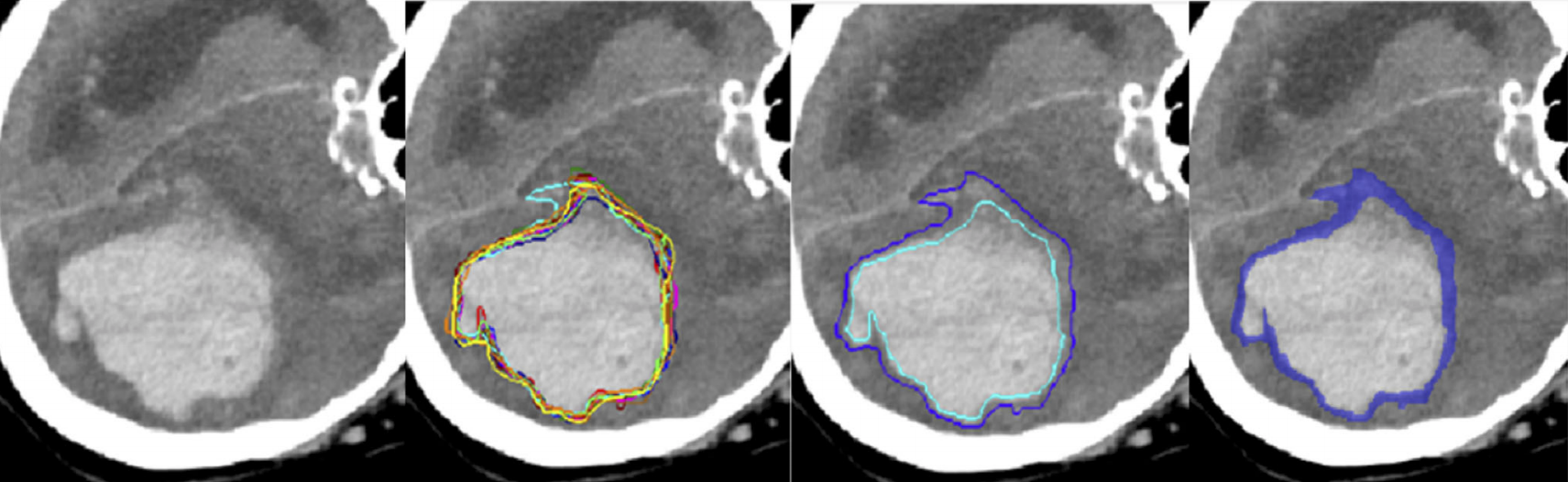
Data¶
Training and test data will comprise 7 binary segmentation tasks in four different CT and MR data sets. Some of the data sets contain more than one binary segmentation (sub-)task, e.g., different sub-structures of tumor or anatomy need to be segmented.
All data sets have about 50 to 100 cases featuring one selected 2D slice each. Each structure of interest is segmented between three and seven times by different experts, and individual segmentations are made available. The task is to delineate structures in a given slice, and to match the distribution – or spread – of the expert's annotations well.
The data is split accordingly in four different image sets, and for each case in those sets, binary labels are given for each segmentation task.
The following data and tasks are available:
- Prostate images (MRI): 55 cases, two segmentation tasks, six
annotations (except one subject has only 5 annotations);
- Brain growth images (MRI): 39 cases, one segmentation task, seven
annotations;
- Brain tumor images (multimodal MRI): 32 cases, three segmentations
tasks, three annotations [Please note: this data set will receive the
additional case in the near future];
- Kidney images (CT): 24 cases, one segmentation task, three
annotations;
Data is available as .nii files with 2D slices. For “Brain tumor” the file contains slices of all four MR modalities.
Metric and algorithmic comparison¶
Each participant will have to segment the given binary structures and predict the distribution of the experts' labels by returning one mask with continuous values in between 0 and 1 that is supposed to reproduce the cumulated segmentations of the experts.
Predictions and continuous ground truth labels are compared by thresholding the continuous labels at predefined thresholds and calculating the volumetric overlap of the resulting binary volumes using Dice score (the** continuous** ground truth labels are obtained by averaging multiple experts annotations). To this end, ground truth and prediction are binarized at ten probability levels (that are 0.1, 0.2, ..., 0.8, 0.9). Dice scores for all thresholds will be averaged.
Dice scores will be averaged across all tasks and all image data sets. The participant performing best according to this average will be named the "winner" of the challenge. Dice scores have many shortcomings, in particular in the novel application domain of uncertainty-aware image quantification. To this end, additional metrics will be calculated and reported for the final analysis, and alternative ranking schemes will be evaluated and presented in the final challenge summary paper.
Participation only in a single task is allowed (just submit the results for the task of your choice), but the winner is determined based on the average score over all the tasks.
It is not mandatory to have a single algorithm for all 7 tasks.
Results of all participants with viable solutions will be reported in a summary paper that will be made available on arxiv immediately after MICCAI conference.
Participation¶
Participants register via "Join" button.
Training and validation data with annotations are available from [here] (password: qubic1917), and test data will be made available to registered participants in early September. The training data is featuring a subset of cases (labeled as ‘evaluation’ set) that can be used for leaderboard evaluation in the online submission system. This data has labels available and is to be used for training the final algorithm as well.
For evaluation, test sets will be made available to the participants at a time point, and they have to upload segmentations within 48 hours. For each task, the number of test cases corresponds to about 20% of the number of training cases.
Update (15/07): validation set are released!
Update (21/08): new annotations from additional raters for brain tumor task are released!
Update (11/09): test set will be released on 16/09.
Update (16/09): test phase started!
Submission data structure¶
As test data, participants will receive images without annotations for all tasks. Participants are encouraged to submit segmentations (i.e. probablity maps) for all 7 tasks (3 for brain tumor, 2 for prostate, 1 for brain growth and 1 for the kidney dataset). The submission folder should be zipped and follow the structure and naming convention of the training data folders. E.g. the tree of the submission folder (results.zip) will be like:
Results --- brain-tumor --- cases*
| task01.nii.gz
| task02.nii.gz
| task03.nii.gz
brain-growth --- cases*
| task01.nii.gz
prostate --- cases*
| task01.nii.gz
| task02.nii.gz
kidney --- cases*
| task01.nii.gz
Timeline¶
- End of June: training data available, registration is open
- Mid of July: validation data available, the leaderboard is open (on the "leaderboard" the predictions are compared with validation ground truth data, so the upload serves only verification of the submission format. When we release the test data the submission format will be identical.)
- September 16th: test data will be made available to the groups, segmentations need to be uploaded. Please contact orgainzers regarding the details.
- October 8th at MICCAI: results will be announced
- October after MICCAI: arxiv paper will be uploaded
The best prizes ever¶
All participants with viable submissions will become co-authors in the challenge summary paper that is to be released on Arxiv directly after the 2020 challenge and will be submitted to a community journal, such as Medical Image Analysis or IEEE Transactions of Medical Imaging after the 2021 edition of QUBIQ.
Contacts¶
If you have any questions, please contact qubiq.miccai@gmail.com or hongwei.li@tum.de