Challenge background¶
The Qubiq segmentation challenge aims at quantifying uncertainty in medical image quantification tasks.
Motivation¶
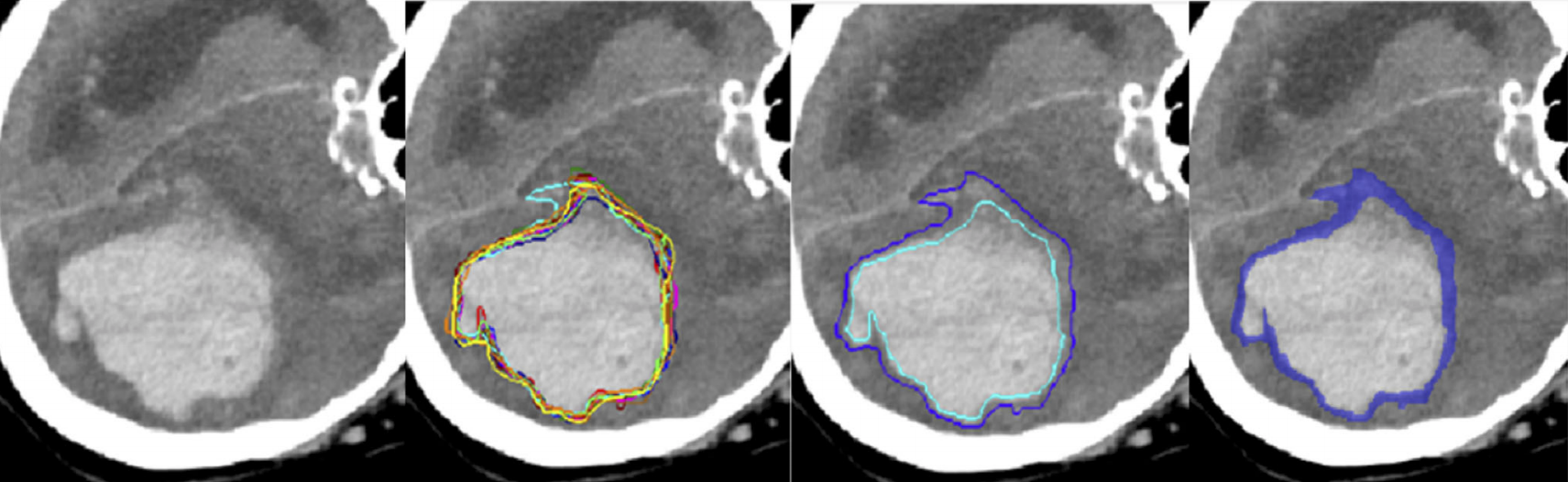
A study on inter-observer variability of manual contour delineation in CT was carried out by L. Joskowicz *et al. * [1] and another multi-reader study in a different MRI segmentation task by E. Konukoglu and colleagues [2]. The objective of both studies was to quantify the interobserver variability of manual delineation of lesions and organ contours in clinical images to establish a reference standard for clinical decision making and for the evaluation of automatic segmentation algorithms. It was observed that the variability in manual delineations for different structures and observers is large and spans a wide range across a variety of structures and pathologies. This variability (that is a property of the biological problem, the imaging modality, and the expert annotators) is – as of now - not sufficiently considered in the design of computerized algorithms for medical image quantification. So far, uncertainties in predicted image segmentation are derived from general considerations of the statistical model, from resampling training data sets in ensemble approaches, or from systematic modifications of the predictive algorithm as in ‘drop-out’ procedures of deep learning procedures. At the same time, the definition of when the outline of an image structure to be quantified is ‘uncertain’ is a task- and data-dependent property of the quantification that can – and maybe has to – be directly inferred from human expert annotations. So far, there are no data sets available for evaluating the accuracy of probabilistic model predictions against such expert generated truth and there is no consensus on what procedures for uncertainty quantification return realistic estimates, and what procedures do not.
Purpose¶
The purpose of the challenge is to benchmark segmentation algorithms returning uncertainty estimates (probability scores, variability regions, etc.) of structures in medical imaging segmentation tasks. Specifically, the algorithmic output will be compared against uncertainties that human annotators attribute to the local delineation of various image structures of diagnostic relevance, such as lesions or anatomical features. Structures in several CT and MR image data sets have repeatedly been annotated by a group of experts to quantify the variability of boundary delineations. Tasks include the segmentation of lesions, such as brain tumors, as well as anatomical structures in kidneys, prostates, or the prenatal brain.
Organizers¶
Bjoern Menze, Technical University of Munich
Leo Joskowicz, The Hebrew University of Jerusalem
Spyridon Bakas, University of Pennsylvania
Andras Jakab, University of Zürich
Ender Konukoglu, ETH Zürich
Anton Becker, MSKCC & Unispital Zürich
Supporting Team¶
Amir Bayat, Technical University of Munich
Hongwei (Bran) Li, Technical University of Munich
Dhritiman Das, Technical University of Munich
Johannes Paetzold, Technical University of Munich
Christoph Berger, Technical University of Munich
Fernando Navarro, Technical University of Munich
Florian Kofler, Technical University of Munich
Giles Titteh, Technical University of Munich
Ivan Ezhov, Technical University of Munich
Kelly Payette , Kinderspital Zürich
If you have any questions, please contact: qubiq.miccai@gmail.com or hongwei.li@tum.de